dixonfix - A tool for correcting fat-water swaps in Dixon MRI
Software
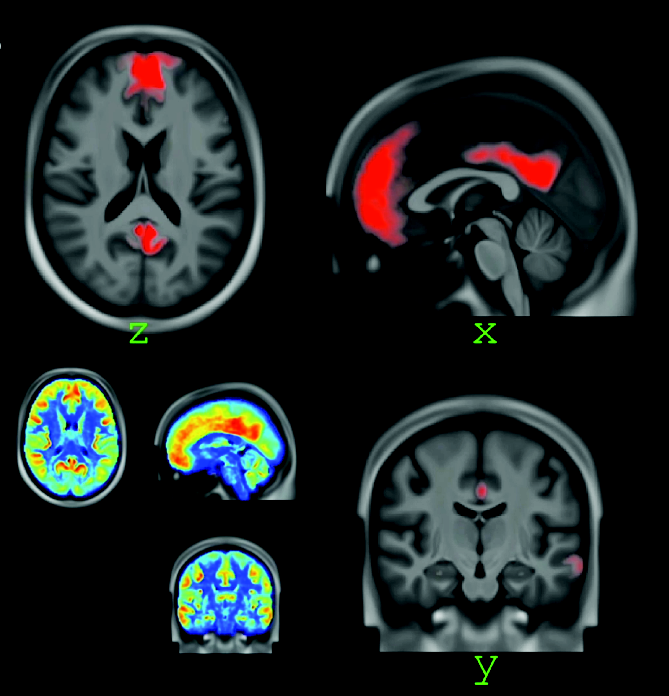
An activation-based visual attribution method for irregular graphs, which works integrated with graph convolutional neural networks (GCNs).
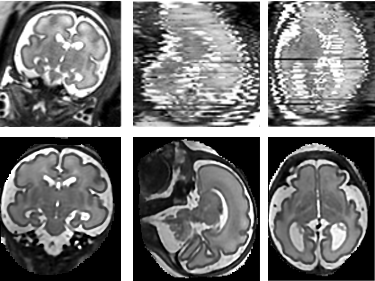
Moving objects cause motion artefacts when their enclosing volume is acquired as a stack of image slices. We present a fast multi-GPU accelerated implementation of slice-to-volume registration based super-resolution reconstruction with automatic outlier rejection and intensity bias correction.
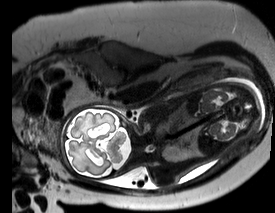
This tool implements a novel method for the correction of motion artifacts as acquired in fetal Magnetic Resonance Imaging (MRI) scans of the whole uterus.
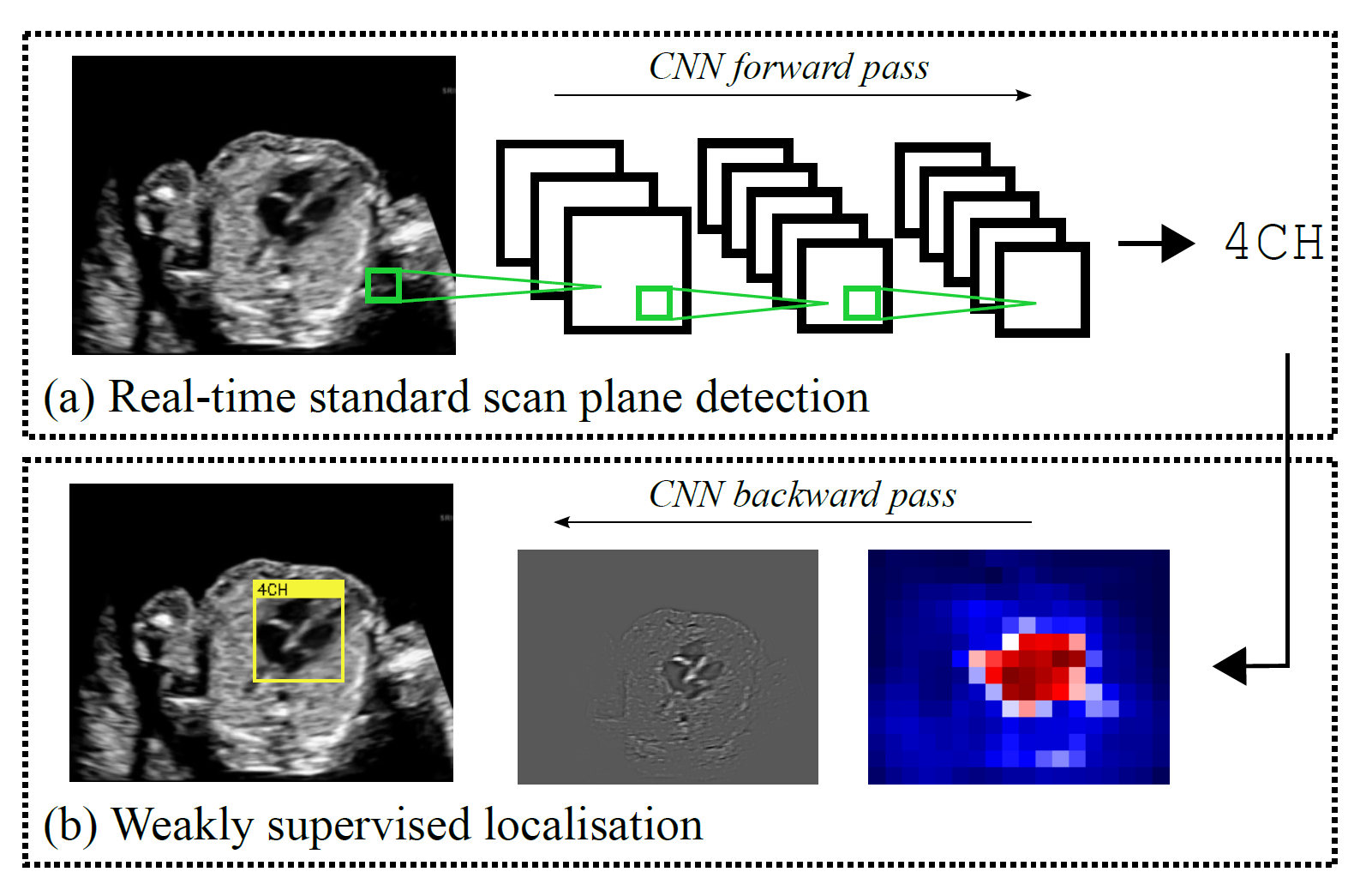
we propose a novel method based on convolutional neural networks which can automatically detect 13 fetal standard views in freehand 2D ultrasound data as well as provide a localisation of the fetal structures via a bounding box.

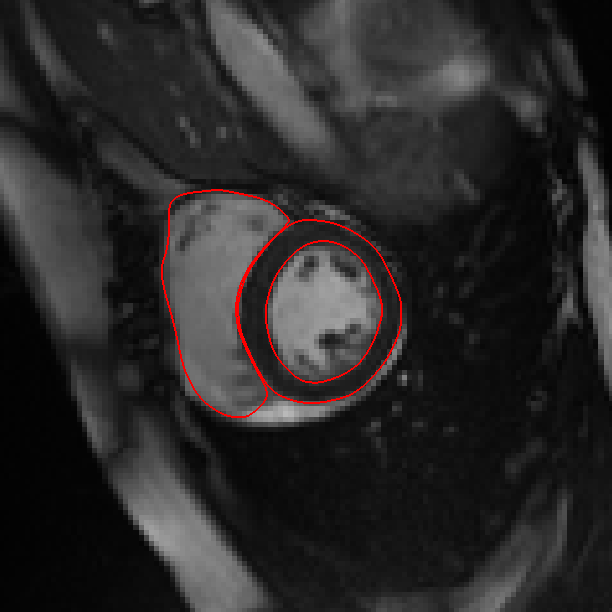
CIMAS is a pipeline for cardiac MR image segmentation. It has been successfully applied to clinical research, segmenting data from the UK Digital Heart project and the UK Biobank project.
OpenMOLE is a workflow engine for executing naturally parallel processes on massively parallel environments. It is free and open-source.

Software which performs whole-brain segmentation of a T1-weighted magnetic resonance brain image. The MALP-EM pipeline includes bias correction, brain extraction, label propagation using multiple atlases, label fusion and finally label refinement using the EM algorithm.
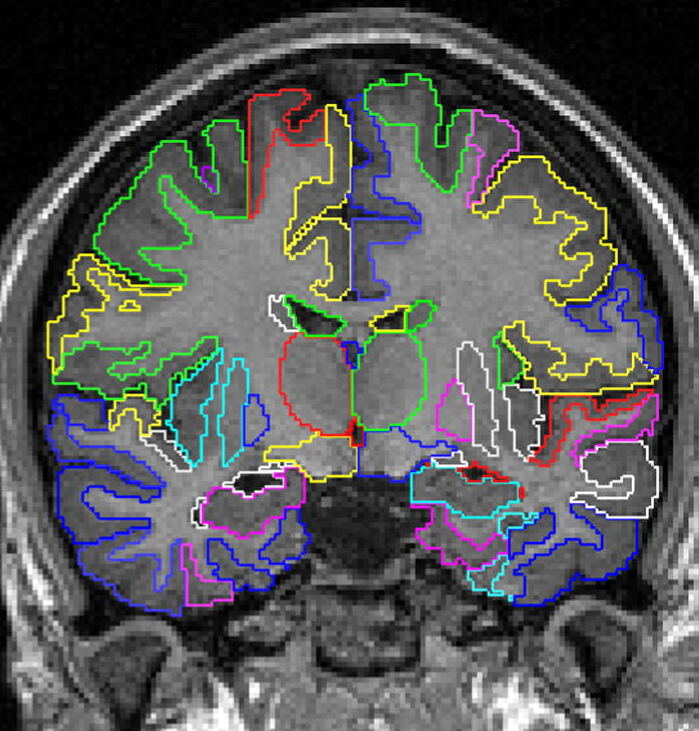
The Developing Brain Region Annotation With Expectation-Maximization software segments neonatal brain MR images into multiple labels.
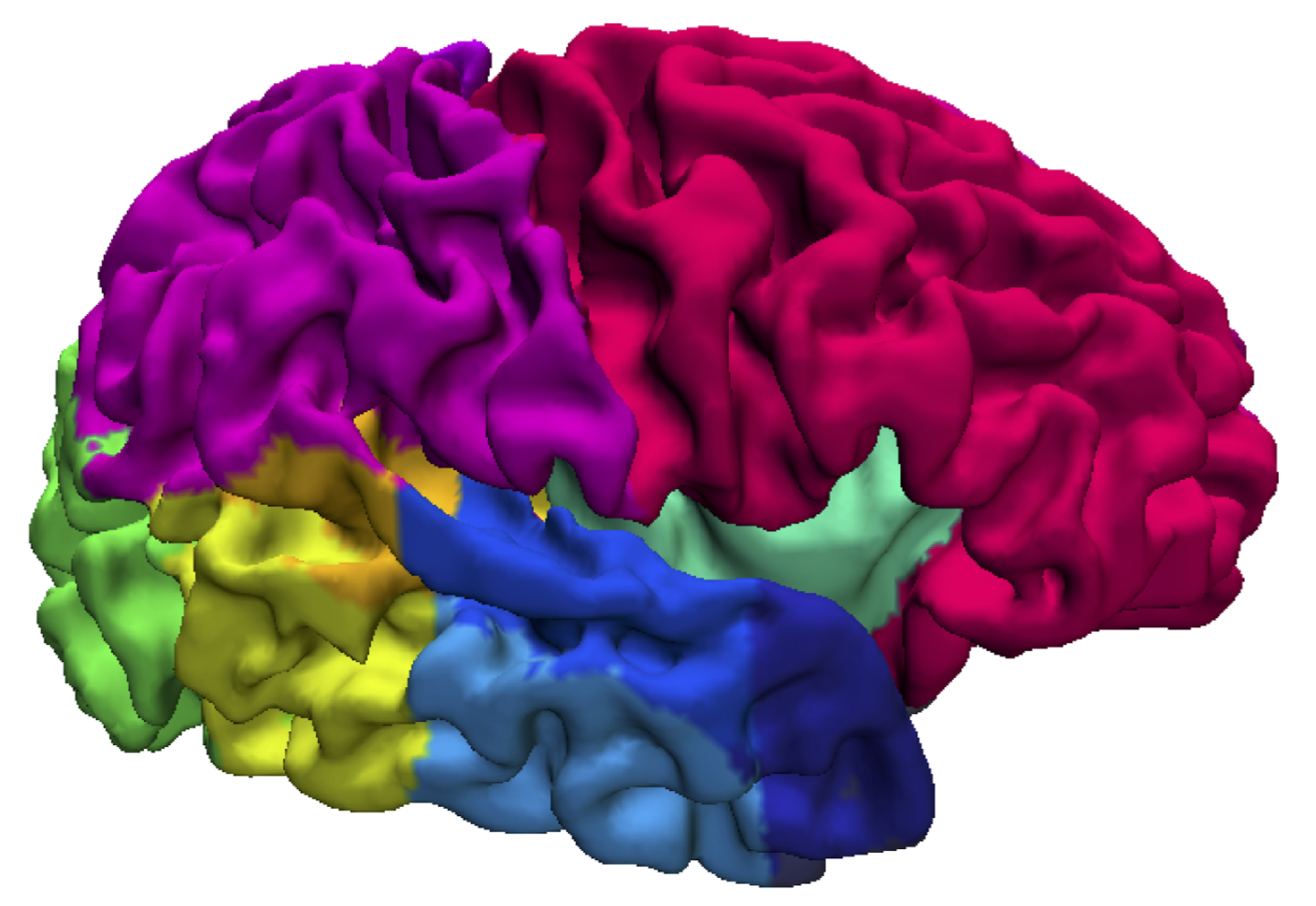
DeepMedic is our software for brain lesion segmentation based on a multi-scale 3D Deep Convolutional Neural Network coupled with a 3D fully connected Conditional Random Field.
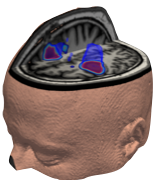
The Medical Image Registration ToolKit provides a collection of libraries and commands to assist in processing and analyzing medical imaging data, including rigid, affine, and deformable registration.
Matlab implementation of the joint spectral decomposition approach for cortical parcellation of the human brain using rs-fMRI.
Matlab implementation of the supervertex clustering algorithm.

Implementation of Neighbourhood Approximation Forests and Weighted Spectral Distance
Real-time dense visual SLAM system capable of capturing comprehensive dense globally consistent surfel-based maps of room scale environments explored using an RGB-D camera.
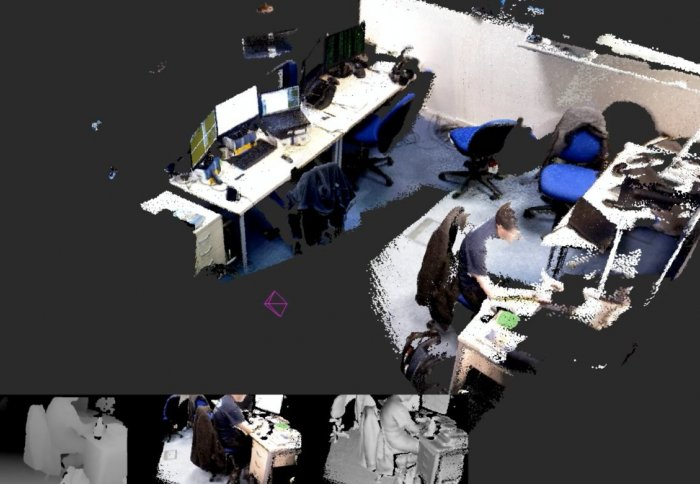

Image registration tool using discrete optimization
also available through Anaconda Python
With examples for temporal sparse free form deformation.
Patch-based Evaluation of Image Segmentation
A Python library for patch-based segmentation with spatial context.
A collection of fast max flow optimization methods for image segmentation.
Software for medical image registration including rigid, affine and non-rigid registration.

